Advanced Light Microscopy

Overview
Light microscopy is a powerful technique that allows scientists to magnify and study tiny structures that are invisible to the naked eye, such as cells and molecules.
By using visible light and specialized lenses, researchers can observe the intricate details of biological samples in real time.
At IFOM, the Advanced Light Microscopy Unit provides cutting-edge imaging technologies that allow scientists to explore the microscopic world in unprecedented detail.
We offer high-resolution and super-resolution microscopy techniques, which go beyond the limits of conventional light microscopes.
These advanced methods enable scientists to visualize structures down to the nanometer scale—helping to unravel key processes in cell biology, cancer research, and beyond.
Our team of experts supports scientists at every stage, from preparing and imaging samples to analyzing complex data.
Over the years, we have built a state-of-the-art facility specifically designed to advance cancer research.
By combining sophisticated imaging technologies with specialized expertise, we aim to better understand cancer at different levels—from single molecules to entire tissues—ultimately contributing to new discoveries and potential treatments.
Advanced Microscopy Techniques
To enhance our understanding of cancer biology at multiple scales, we have dedicated substantial efforts to optimizing and expanding access to a broad range of optical microscopy techniques and applications, including:
- Molecular Mobility:
Optogenetics, Photoactivation/Photoswitching, Fluorescence Loss in Photobleaching (FLIP), Fluorescence Recovery After Photobleaching (FRAP) - Microinjection/Micromanipulation
- Non-Linear Microscopy:
Two-Photon Excitation (2PE), Second Harmonic Generation (SHG) - Protein Interactions:
Colocalization, Fluorescence Resonance Energy Transfer (FRET), Fluorescence Lifetime Imaging - Laser-Induced Stimulation:
DNA Damage Response, Laser Ablation - Mechanobiology
- Nanoscopy:
Super-Resolution Microscopy (STED, STORM, PALM) - Correlative Light Electron Microscopy (CLEM)
Sample Preparation Support
We provide guidance on sample preparation and staining, for both live-cell and fixed-cell experiments. In particular, we offer:
- protocols for immunofluorescence and live-cell staining
- protocols for clarification of organoids and tissues
- Reagent supplies (commonly used antibodies, Dapi for nuclei staining, conjugated antibodies for STED microscopy)
- Troubleshooting for improved sample preparation and acquisition
Image Analysis & Data Processing
With approximately 180 custom analysis protocols developed annually, our team provides support immediately after the collection of the data, from image processing to data analysis and interpretation. All the plugins and pipelines can be developed in open-source software (Fiji, CellProfiler, QuPath) or commercial platforms (Matlab, Arivis, Harmony, Prism). Examples of implemented tools include:
- Cell, Nuclei, and Particle Detection:
Segmentation and classification using classical methods or machine learning tools (2D and 3D). - Cell and Particle Tracking:
Pipelines for single-cell tracking, Particle Image Velocimetry (PIV), Wound Healing Assays, and particle trafficking. - Molecular Mobility:
FRAP analysis. - Label-free imaging analysis:
Machine learning tools for collagen fiber orientation analysis. - Protein Interactions:
Quantification of colocalization, foci formation (2D and 3D), and FRET analysis. - Immunohistochemistry imaging analysis:
Object detection pipelines using color threshold segmentation or machine learning (pre-trained or custom-trained models), applicable to whole-slide and single-image analyses. - AI-Assisted Image Analysis
Integration of artificial intelligence and machine learning tools for enhanced segmentation, tracking, and quantitative microscopy workflows.
Training & Support
We provide comprehensive support tailored to both senior researchers and early-career scientists, offering personalized training to foster their professional development. Our team ensures that users gain the necessary expertise to fully exploit the advanced microscopy resources available at IFOM.
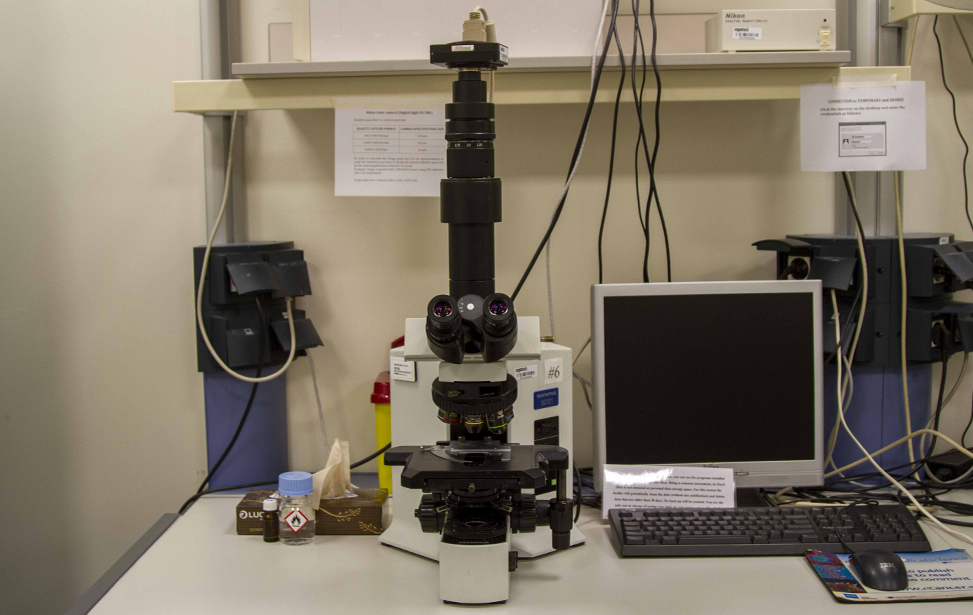
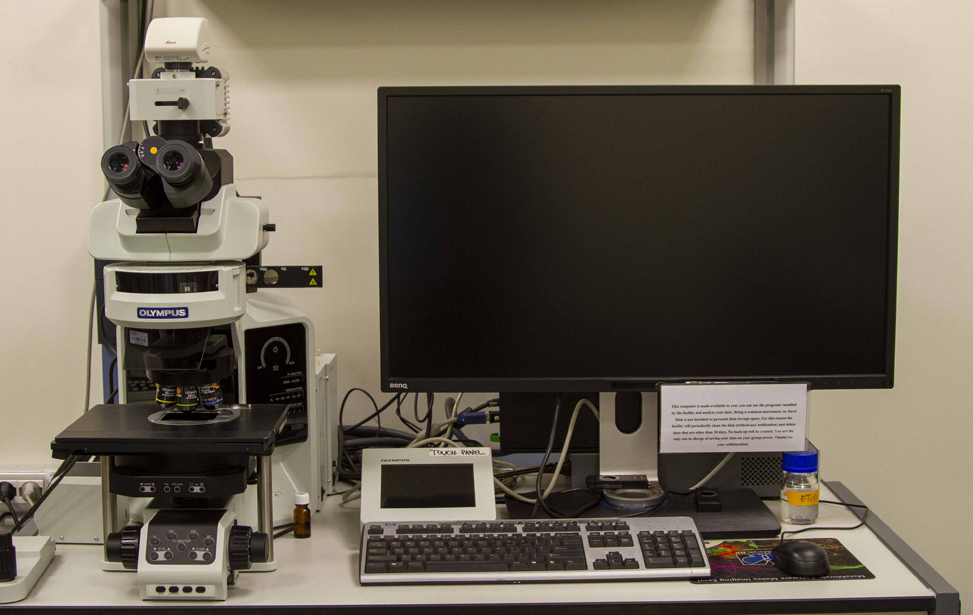
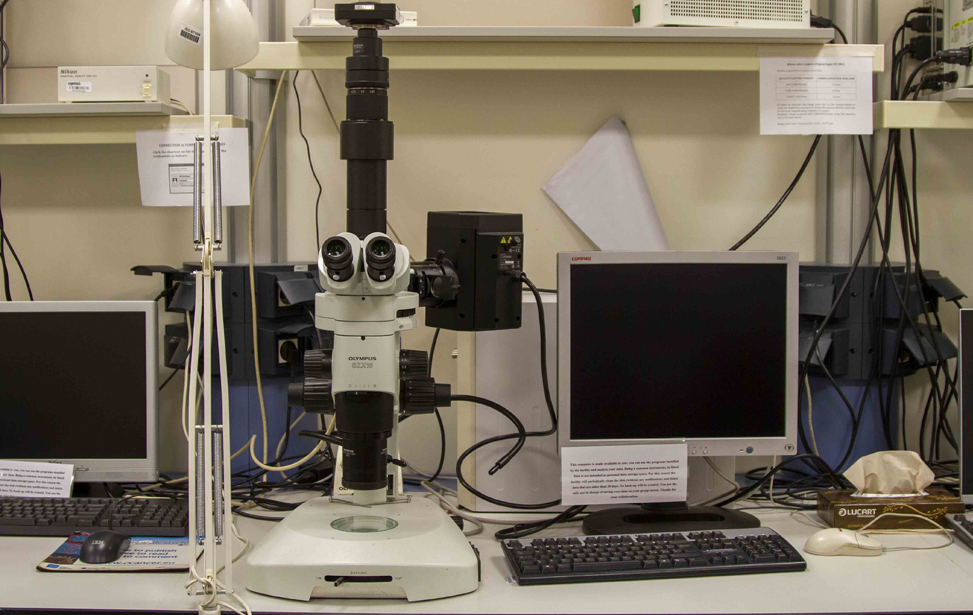
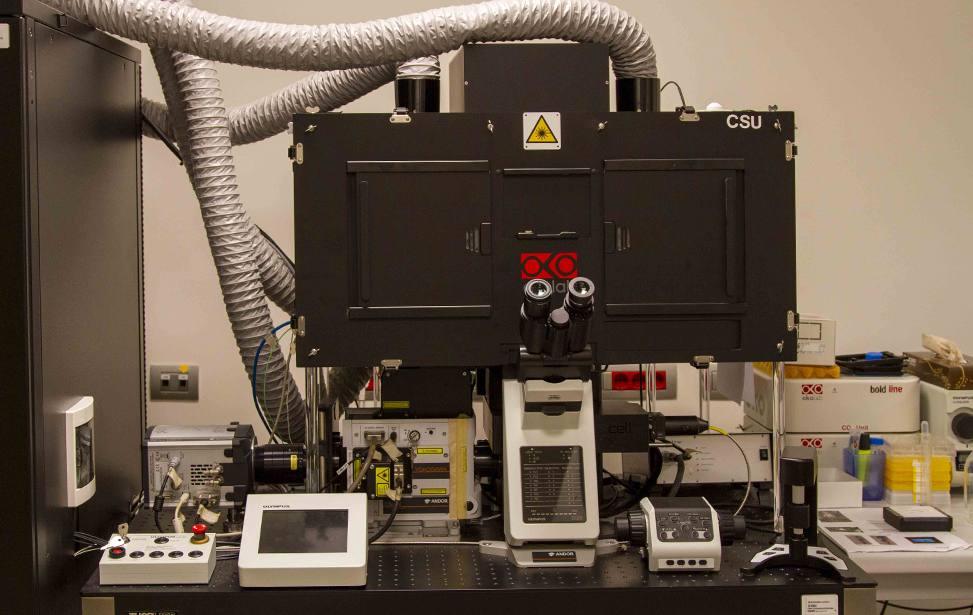
Available Microscopes
- Widefield microscopes(4 upright, 1 inverted, 1 stereo) for fluorescence and/or transmitted light microscopy
- IXplore (Evident): Widefield Microscope for live-cell imaging or fixed samples
- Thunder (Leica): Widefield Microscope with computational clearing for live-cell imaging or fixed samples
- DeltaVision (GE Healthcare): Wide Field Microscope with Deconvolution for live-cell imaging or fixed samples
- TIRFM(Leica): Total internal reflection fluorescence microscope, for live-cell imaging or fixed samples.
- Spinning Disk CSU (Evident): Widefield Microscopy, spinning-disk confocal Microscopy, Photomanipulation for live-cell imaging or fixed samples
- TCS SP2 AOBS (Leica): Confocal Microscopy
- TCS SP5 Inverted (Leica): Confocal Microscopy, non-linear microscopy, Photo manipulation for fixed samples or live cell imaging
- TCS SP5 Upright (Leica):Confocal Microscopy, non-linear microscopy for fixed samples or live cell imaging, Intravital microscopy
- TCS SP8 STED (Leica): Confocal Microscopy, Super Resolution microscopy, FLIM (Fluorescence Lifetime Imaging Microscopy), Photomanipulation, for fixed samples or live-cell imaging
- TCS SP8 DLS (Leica): Confocal Microscopy for fixed samples, Digital Light-Sheet microscopy for live-cell imaging or fixed samples
- Stellaris 8 DIVE (Leica): Confocal Microscopy, non-linear microscopy, FLIM, Photomanipulation, for live-cell imaging or fixed samples, Intravital microscopy
More information
Other equipment
- Envision (Revvity): Micro Plate reader
- AIS2: Microinjection System
- CellASIC ONIX2 (Merck): Microfluidic System
Facility Acknowledgment
According to our Core Facilities Guidelines, if Facility services were used in work described in a publication, the Facility must be acknowledged, citing both the Facillity name and its Research Resource Identifier (RRID). A typically acknowledgement might look like:
The research described was supported by the IFOM Advanced Light Microscopy Core Facility (RRID: SCR_026866)
Co‑Authorship
If a staff member of an IFOM or Cogentech Core Facility contributes substantially to a published work, they should be included as a co‑author.
More information is available in Recommended guidelines and Acknowledging and citing core facilities
Examples of substantial contributions that may justify co‑authorship include:
- Contribution to experimental design or optimization
- Development or significant adaptation of methods or protocols
- Advanced data analysis and interpretation
- Intellectual input influencing the scientific direction of the study
Routine technical support or standard service provision generally warrants acknowledgment rather than co‑authorship.
- PMID: 39536107
USP1 deubiquitinates PARP1 to regulate its trapping and PARylation activity.
Nespolo A, Stefenatti L, Pellarin I, Gambelli A, Rampioni Vinciguerra GL, Karimbayli J, Barozzi S, Orsenigo F, Spizzo R, Nicoloso MS, Segatto I, D'Andrea S, Bartoletti M, Lucia E, Giorda G, Canzonieri V, Puglisi F, Belletti B, Schiappacassi M, Baldassarre G, Sonego M.
2024 Jan 2;84(1):133-153. doi: 10.1158/0008-5472.CAN-23-1129. PMID: 37855660 Free PMC article. - PMID: 39218980
Emerin mislocalization during chromatin bridge resolution can drive prostate cancer cell invasiveness in a collagen-rich microenvironment.
Popęda M, Kowalski K, Wenta T, Beznoussenko GV, Rychłowski M, Mironov A, Lavagnino Z, Barozzi S, Richert J, Bertolio R, Myszczyński K, Szade J, Bieńkowski M, Miszewski K, Matuszewski M, Żaczek AJ, Braga L, Del Sal G, Bednarz-Knoll N, Maiuri P, Nastały P.
Exp Mol Med. 2024 Sep;56(9):2016-2032. doi: 10.1038/s12276-024-01308-w. Epub 2024 Sep 2. PMID: 39218980 Free PMC article. - PMID: 39155303
CDK12 controls transcription at damaged genes and prevents MYC-induced transcription-replication conflicts.
Curti L, Rohban S, Bianchi N, Croci O, Andronache A, Barozzi S, Mattioli M, Ricci F, Pastori E, Sberna S, Bellotti S, Accialini A, Ballarino R, Crosetto N, Wade M, Parazzoli D, Campaner S.
Nat Commun. 2024 Aug 18;15(1):7100. doi: 10.1038/s41467-024-51229-5.PMID: 39155303 Free PMC article. - PMID: 39208097
A p62-dependent rheostat dictates micronuclei catastrophe and chromosome rearrangements.
Martin S, Scorzoni S, Cordone S, Mazzagatti A, Beznoussenko GV, Gunn AL, Di Bona M, Eliezer Y, Leor G, Ben-Yishay T, Loffreda A, Cancila V, Rainone MC, Ippolito MR, Martis V, Bedin F, Garrè M, Vaites LP, Vasapolli P, Polo S, Parazzoli D, Tripodo C, Mironov AA, Cuomo A, Ben-David U, Bakhoum SF, Hatch EM, Ly P, Santaguida S.
Science. 2024 Aug 30;385(6712):eadj7446. doi: 10.1126/science.adj7446. Epub 2024 Aug 30.PMID: 39208097 Free PMC article. - PMID: 38977673
Mechanically induced topological transition of spectrin regulates its distribution in the mammalian cell cortex.
Ghisleni A, Bonilla-Quintana M, Crestani M, Lavagnino Z, Galli C, Rangamani P, Gauthier NC.
Nat Commun. 2024 Jul 8;15(1):5711. doi: 10.1038/s41467-024-49906-6.PMID: 38977673 Free PMC article. - PMID: 38990945
Mechanical stress during confined migration causes aberrant mitoses and c-MYC amplification.
Bastianello G, Kidiyoor GR, Lowndes C, Li Q, Bonnal R, Godwin J, Iannelli F, Drufuca L, Bason R, Orsenigo F, Parazzoli D, Pavani M, Cancila V, Piccolo S, Scita G, Ciliberto A, Tripodo C, Pagani M, Foiani M.
Proc Natl Acad Sci U S A. 2024 Jul 16;121(29):e2404551121. doi: 10.1073/pnas.2404551121. Epub 2024 Jul 11.PMID: 38990945 Free PMC article. - PMID: 39087467
GPR126 is a specifier of blood-brain barrier formation in the mouse central nervous system.
Kakogiannos N, Scalise AA, Martini E, Maderna C, Benvenuto AF, D'Antonio M, Carmignani L, Magni S, Gullotta GS, Lampugnani MG, Iannelli F, Beznoussenko GV, Mironov AA, Cerutti C, Bentley K, Philippides A, Zanardi F, Bacigaluppi M, Sigismund S, Bassani C, Farina C, Martino G, De Giovanni M, Dejana E, Iannacone M, Inverso D, Giannotta M.
J Clin Invest. 2024 Jun 6;134(15):e165368. doi: 10.1172/JCI165368.PMID: 39087467 Free PMC article. -
PMID: 38088930
Cell stretching activates an ATM mechano-transduction pathway that remodels cytoskeleton and chromatin.
Bastianello G, Porcella G, Beznoussenko GV, Kidiyoor G, Ascione F, Li Q, Cattaneo A, Matafora V, Disanza A, Quarto M, Mironov AA, Oldani A, Barozzi S, Bachi A, Costanzo V, Scita G, Foiani M.
Cell Rep. 2023 Dec 26;42(12):113555. doi: 10.1016/j.celrep.2023.113555. Epub 2023 Dec 11.PMID: 38088930 Free PMC article. -
PMID: 37925537
Alternative lengthening of telomeres (ALT) cells viability is dependent on C-rich telomeric RNAs.
Rosso I, Jones-Weinert C, Rossiello F, Cabrini M, Brambillasca S, Munoz-Sagredo L, Lavagnino Z, Martini E, Tedone E, Garre' M, Aguado J, Parazzoli D, Mione M, Shay JW, Mercurio C, d'Adda di Fagagna F.
Nat Commun. 2023 Nov 4;14(1):7086. doi: 10.1038/s41467-023-42831-0. PMID: 37925537 Free PMC article. -
PMID: 37713487
Clonal cooperation through soluble metabolite exchange facilitates metastatic outgrowth by modulating Allee effect.
Hershey BJ, Barozzi S, Orsenigo F, Pompei S, Iannelli F, Kamrad S, Matafora V, Pisati F, Calabrese L, Fragale G, Salvadori G, Martini E, Totaro MG, Magni S, Guan R, Parazzoli D, Maiuri P, Bachi A, Patil KR, Cosentino Lagomarsino M, Havas KM.
Sci Adv. 2023 Sep 15;9(37):eadh4184. doi: 10.1126/sciadv.adh4184. Epub 2023 Sep 15. PMID: 37713487 Free PMC article. -
PMID: 36581770
Tissue fluidification promotes a cGAS-STING cytosolic DNA response in invasive breast cancer.
Frittoli E, Palamidessi A, Iannelli F, Zanardi F, Villa S, Barzaghi L, Abdo H, Cancila V, Beznoussenko GV, Della Chiara G, Pagani M, Malinverno C, Bhattacharya D, Pisati F, Yu W, Galimberti V, Bonizzi G, Martini E, Mironov AA, Gioia U, Ascione F, Li Q, Havas K, Magni S, Lavagnino Z, Pennacchio FA, Maiuri P, Caponi S, Mattarelli M, Martino S, d'Adda di Fagagna F, Rossi C, Lucioni M, Tancredi R, Pedrazzoli P, Vecchione A, Petrini C, Ferrari F, Lanzuolo C, Bertalot G, Nader G, Foiani M, Piel M, Cerbino R, Giavazzi F, Tripodo C, Scita G.
Nat Mater. 2023 May;22(5):644-655. doi: 10.1038/s41563-022-01431-x. Epub 2022 Dec 29.PMID: 36581770 Free PMC article. -
PMID: 36894671
SARS-CoV-2 infection induces DNA damage, through CHK1 degradation and impaired 53BP1 recruitment, and cellular senescence.
Gioia U, Tavella S, Martínez-Orellana P, Cicio G, Colliva A, Ceccon M, Cabrini M, Henriques AC, Fumagalli V, Paldino A, Presot E, Rajasekharan S, Iacomino N, Pisati F, Matti V, Sepe S, Conte MI, Barozzi S, Lavagnino Z, Carletti T, Volpe MC, Cavalcante P, Iannacone M, Rampazzo C, Bussani R, Tripodo C, Zacchigna S, Marcello A, d'Adda di Fagagna F.
Nat Cell Biol. 2023 Apr;25(4):550-564. doi: 10.1038/s41556-023-01096-x. Epub 2023 Mar 9.PMID: 36894671 Free PMC article. -
PMID: 36460033
Decellularized extracellular matrix as scaffold for cancer organoid cultures of colorectal peritoneal metastases.
Varinelli L, Guaglio M, Brich S, Zanutto S, Belfiore A, Zanardi F, Iannelli F, Oldani A, Costa E, Chighizola M, Lorenc E, Minardi SP, Fortuzzi S, Filugelli M, Garzone G, Pisati F, Vecchi M, Pruneri G, Kusamura S, Baratti D, Cattaneo L, Parazzoli D, Podestà A, Milione M, Deraco M, Pierotti MA, Gariboldi M.
J Mol Cell Biol. 2023 Apr 6;14(11):mjac064. doi: 10.1093/jmcb/mjac064. PMID: 36460033 Free PMC article.
Equipment
Gallery
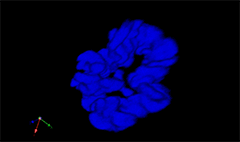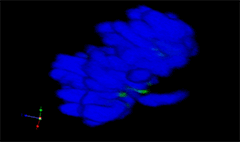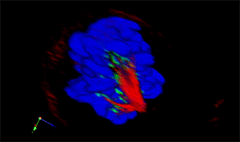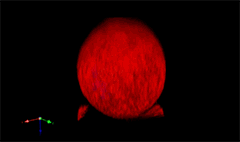
3D rendering of a mitotic He-La cell
(blue chromatin, green CREST, red tubulin)
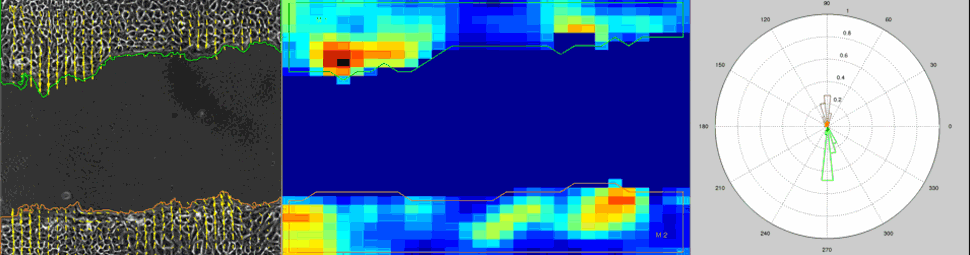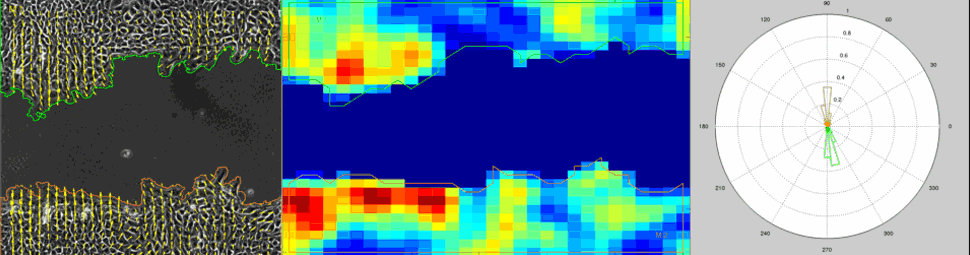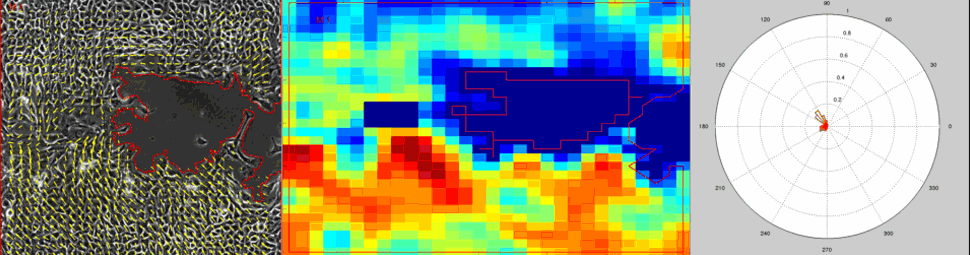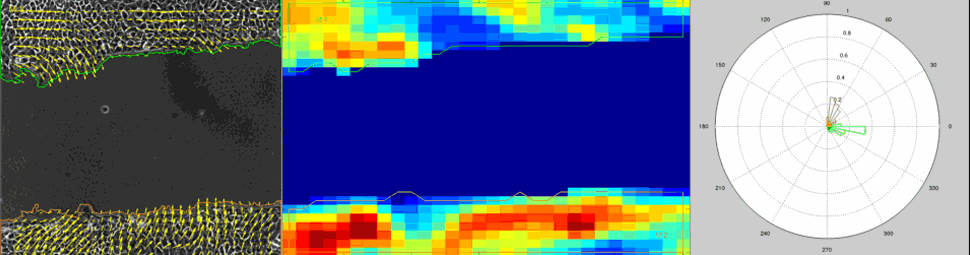
Particle Image Velocimetry (PIV) Analysis of a wound healing experiment

Dario Parazzoli
Dario Parazzoli has a long-standing passion for the intersection of physics and biology. After earning his degree in Physics from the University of Milan, where he studied radionuclide dispersion following the Chernobyl disaster, he began his career in the microscopy industry. For nearly a decade, he worked as a project manager, contributing to advancements in confocal microscopy and high-end solutions for life science imaging.
In 2004, Dario transitioned to biomedical research at the European Institute of Oncology, where he played a key role in adapting Total Internal Reflection Fluorescence (TIRF) microscopy for long-term live-cell imaging. He later established an imaging facility at the National Institute of Molecular Genetics before joining IFOM in 2009.
As the Head of the Advanced Light Microscopy Unit at IFOM, Dario oversees cutting-edge imaging technologies, supporting cancer research by enabling high-resolution visualization of cellular processes. His expertise continues to drive innovation in biomedical imaging, helping researchers uncover new insights into cancer biology.






















