Flow Cytometry

Unit info
Flow cytometry is a powerful technology for analyzing and sorting cells based on their physical and molecular characteristics, providing valuable insights for both research and clinical applications. Since its development in the 1960s, it has become an essential tool in scientific research, offering rapid data acquisition, a wide range of applications, and the ability to explore complex biological systems.
In recent years, flow cytometry has undergone exponential advancements. While early instruments could process only a limited number of parameters, gradual technological improvements have significantly expanded their capabilities. The introduction of full-spectrum technology has been a breakthrough, enabling the simultaneous analysis of over 50 parameters, as demonstrated in published studies.
Beyond its analytical power, flow cytometry plays a crucial role in immunology, where the ability to identify and separate specific cell populations is indispensable. More broadly, it continues to drive discoveries in cellular function, disease mechanisms, and therapeutic development.
The Flow Cytometry Core Facility at IFOM provides cutting-edge technology and state-of-the-art instrumentation, managed by a highly experienced team with decades of expertise. The facility supports a wide range of applications, including immunophenotyping, cell cycle analysis, metabolism studies, apoptosis and cell proliferation assays, fluorescence protein analysis, and the detection of cytokines, chemokines, and intra/extracellular proteins.
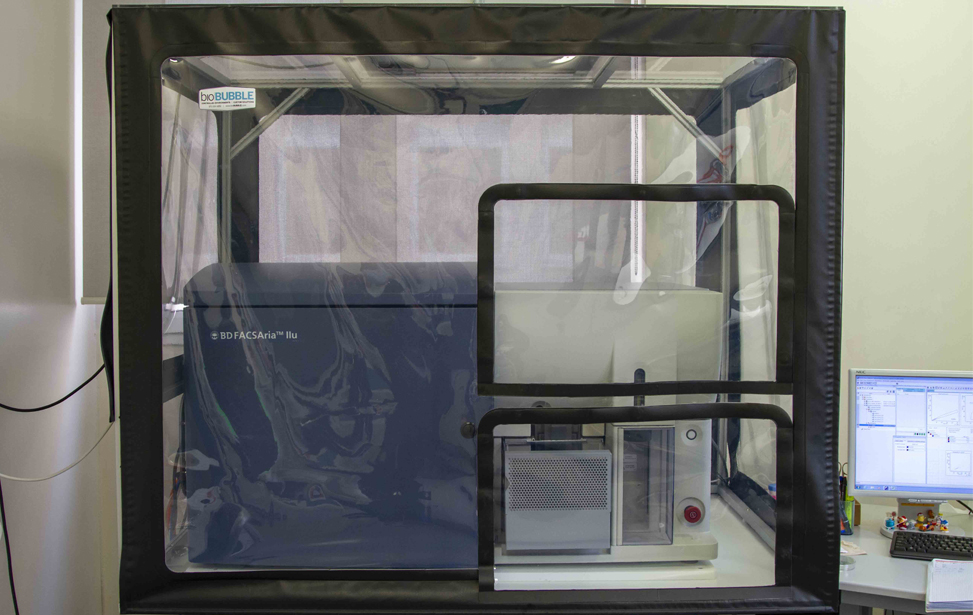
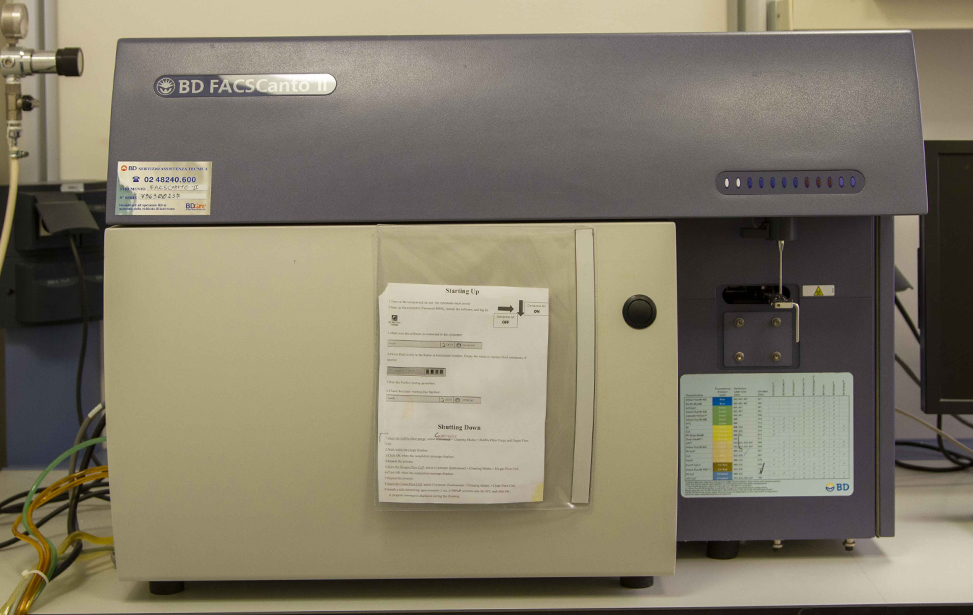
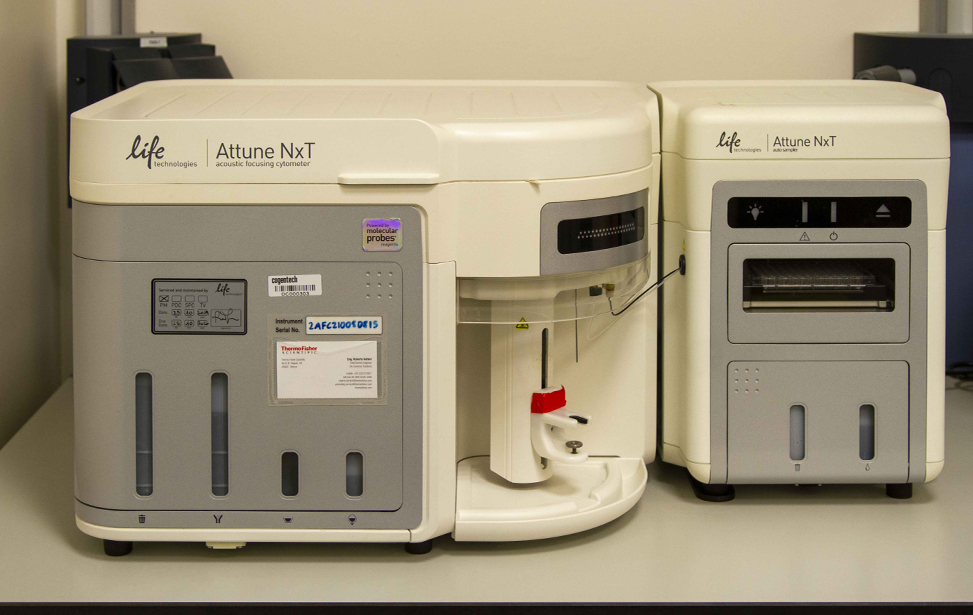
Users have access to both traditional flow cytometers (FACSCanto II, Attune NxT) and advanced spectral instruments (Aurora 5L). Independent use of these instruments is permitted upon completion of a theoretical and practical training course organized by the facility staff.
For cell sorting, access is available only with assistance. The facility offers both conventional sorters (MoFlo Astrios, FACSAria IIU) and a spectral sorter (Aurora CS).
Additionally, the facility collaborates with the bioinformatics team to provide high-parameter data analysis services. Expert staff are also available to offer guidance and support throughout the experimental process, including panel design, optimization, and troubleshooting.
The Flow Cytometry Core Facility ensures optimal instrument setup, maintenance, and performance. The facility staff provides expert guidance on sample preparation, panel design, and gating strategies.
Additionally, the facility maintains close collaborations with industry leaders to test and evaluate cutting-edge technologies and reagents. It also actively engages with other core facilities to share insights, address challenges, and stay updated on advancements in the field.
Data Analysis & Software
The facility supports data analysis using various software platforms, including:
- Kaluza
- FlowJo
- FCS Express
- SpectroFlo
- FACSDiva
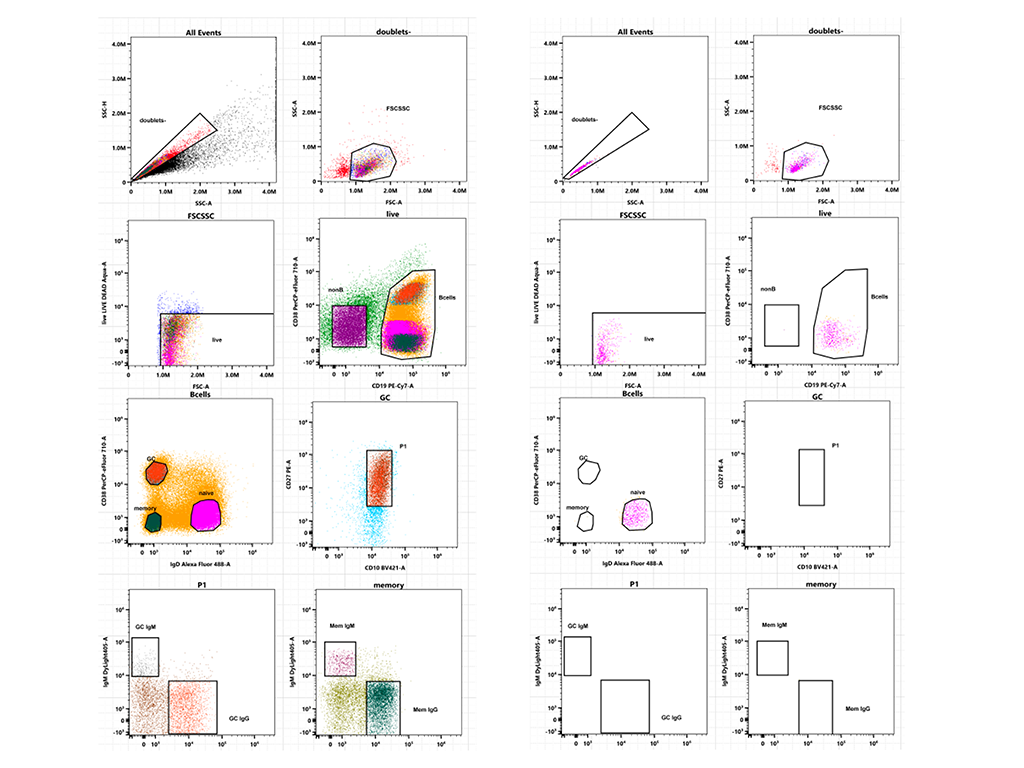
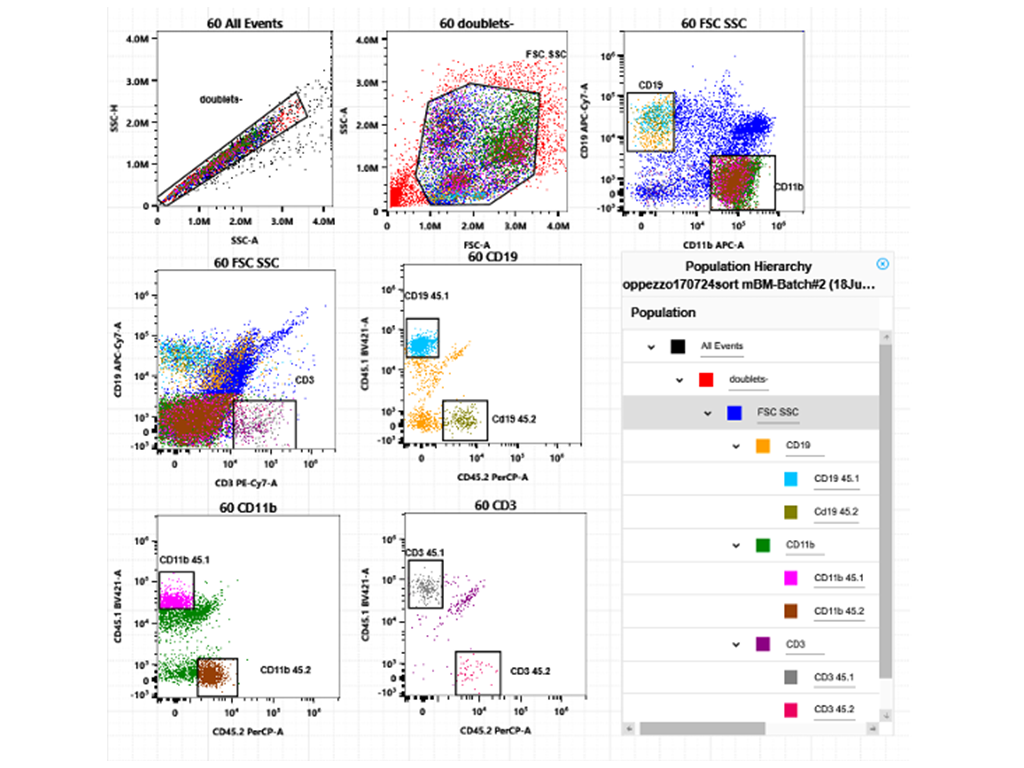
Cell Population Identification
The facility specializes in identifying cell subpopulations from various human and murine samples, tissues, and 3D cultures (spheroids, organoids, co-cultures) using multiparametric staining. These include:
- Immune subpopulations (e.g., Treg, T naive, Tconv, GC B cells, naïve B cells, TIL, TAM)
- Cancer stem cells (e.g., ALDH, CD44/CD24)
- Endothelial cells
- Myeloma cells
- Cardiac fibroblasts
Cell Cycle & Proliferation Analysis
Flow cytometry assays for cell cycle and proliferation in mammalian and yeast cells using:
- PI, Sytox, BrdU, EdU, FUCCI, Cyclins, Ki67
Metabolic Analysis
The facility provides various assays to study cell metabolism, including:
- Glucose uptake (2NDBG)
- Lipid visualization (Bodipy) and lipid droplet analysis (Nile Red)
- Mitochondrial function (JC1, Mitotracker, MitoSOX)
- Intracellular protein detection (TMR HaloTag)
- Reactive oxygen species detection (CellROX, DFCDA)
- ATP/ADP ratio monitoring (Perceval sensor)
Apoptosis & Cell Death Analysis
- Caspase-3, Annexin V/PI, DRAQ7
Fluorescent Protein & Gene Editing Analysis
- Analysis of transduced, transfected, or CRISPR/Cas9-engineered cell lines and primary cells (human and murine)
- Fluorescent protein detection (GFP, RFP, mCherry, Scarlet, Tomato, YFP)
- Lineage tracking via RGB barcoding
- Protein interaction analysis (Duolink flowPLA® kit)
- Endocytosis analysis (Dextran uptake)
- Microplastic uptake analysis in various cell lines and murine tissues
- Antigen quantification per cell (ABC) using QuantiBrite™ beads
Cell Sorting Capabilities
The facility offers sorting for all the above-mentioned cells/populations using:
- Bulk sorting
- Sorting onto specialized supports (glass slides for imaging, 96-well plates for 10x Genomics, sequencing, mass spectrometry, high-content imaging)
- Single-cell sorting (96- or 384-well plates)
- Sorting with specific precautions for metabolomic and NGS analysis
Main Users of the Flowcytometry facility are the IFOM Research Groups
Facility Acknowledgment
According to our Core Facilities Guidelines, if Facility services were used in work described in a publication, the Facility must be acknowledged, citing both the Facillity name and its Research Resource Identifier (RRID). A typically acknowledgement might look like:
The research described was supported by the IFOM Flow Cytometry Core Facility (RRID: SCR_026865)
Co‑Authorship
If a staff member of an IFOM or Cogentech Core Facility contributes substantially to a published work, they should be included as a co‑author.
More information is available in Recommended guidelines and Acknowledging and citing core facilities
Examples of substantial contributions that may justify co‑authorship include:
- Contribution to experimental design or optimization
- Development or significant adaptation of methods or protocols
- Advanced data analysis and interpretation
- Intellectual input influencing the scientific direction of the study
Routine technical support or standard service provision generally warrants acknowledgment rather than co‑authorship.
- PMID: 37713487
Clonal cooperation through soluble metabolite exchange facilitates metastatic outgrowth by modulating Allee effect.
Hershey BJ, Barozzi S, Orsenigo F, Pompei S, Iannelli F, Kamrad S, Matafora V, Pisati F, Calabrese L, Fragale G, Salvadori G, Martini E, Totaro MG, Magni S, Guan R, Parazzoli D, Maiuri P, Bachi A, Patil KR, Cosentino Lagomarsino M, Havas KM. Sci Adv. 2023 Sep 15;9(37):eadh4184. doi: 10.1126/sciadv.adh4184. Epub 2023 Sep 15. Free PMC article. - PMID: 37108076
Short-Term Exposure to Benzo(a)Pyrene Causes Disruption of GnRH Network in Zebrafish Embryos.
Gentile I, Vezzoli V, Martone S, Totaro MG, Bonomi M, Persani L, Marelli F. Int J Mol Sci. 2023 Apr 7;24(8):6913. doi: 10.3390/ijms24086913. Free PMC article. - PMID: 35918044
HDAC6 inhibition decreases leukemic stem cell expansion driven by Hedgehog hyperactivation by restoring primary ciliogenesis.
Pezzotta A, Gentile I, Genovese D, Totaro MG, Battaglia C, Leung AY, Fumagalli M, Parma M, Cazzaniga G, Fazio G, Alcalay M, Marozzi A, Pistocchi A. Pharmacol Res. 2022 Sep;183:106378. doi: 10.1016/j.phrs.2022.106378. Epub 2022 Jul 30. - PMID: 33653693
CD4 T Cell-Dependent Rejection of Beta-2 Microglobulin Null Mismatch Repair-Deficient Tumors.
Germano G, Lu S, Rospo G, Lamba S, Rousseau B, Fanelli S, Stenech D, Le DT, Hays J, Totaro MG, Amodio V, Chilà R, Mondino A, Diaz LA Jr, Di Nicolantonio F, Bardelli A. Cancer Discov. 2021 Jul;11(7):1844-1859. doi: 10.1158/2159-8290.CD-20-0987. Epub 2021 Mar 2. - PMID: 33436662
MITO-Luc/GFP zebrafish model to assess spatial and temporal evolution of cell proliferation in vivo.
de Latouliere L, Manni I, Ferrari L, Pisati F, Totaro MG, Gurtner A, Marra E, Pacello L, Pozzoli O, Aurisicchio L, Capogrossi MC, Deflorian G, Piaggio G. Sci Rep. 2021 Jan 12;11(1):671. PMID: 33436662 Free PMC article. - PMID: 33015043
HDAC8: A Promising Therapeutic Target for Acute Myeloid Leukemia.
Spreafico M, Gruszka AM, Valli D, Mazzola M, Deflorian G, Quintè A, Totaro MG, Battaglia C, Alcalay M, Marozzi A, Pistocchi A. Front Cell Dev Biol. 2020 Sep 4;8:844. doi: 10.3389/fcell.2020.00844. eCollection 2020. PMID: 33015043 Free PMC article.
Pre IFOM Publications – Totaro
- PMID: 33903766
Heme catabolism by tumor-associated macrophages controls metastasis formation.
Consonni FM, Bleve A, Totaro MG, Storto M, Kunderfranco P, Termanini A, Pasqualini F, Alì C, Pandolfo C, Sgambelluri F, Grazia G, Santinami M, Maurichi A, Milione M, Erreni M, Doni A, Fabbri M, Gribaldo L, Rulli E, Soares MP, Torri V, Mortarini R, Anichini A, Sica A. Nat Immunol. 2021 May;22(5):595-606. doi: 10.1038/s41590-021-00921-5. Epub 2021 Apr 26. - PMID: 32265223
Tumor-Derived Prostaglandin E2 Promotes p50 NF-κB-Dependent Differentiation of Monocytic MDSCs.
Porta C, Consonni FM, Morlacchi S, Sangaletti S, Bleve A, Totaro MG, Larghi P, Rimoldi M, Tripodo C, Strauss L, Banfi S, Storto M, Pressiani T, Rimassa L, Tartari S, Ippolito A, Doni A, Soldà G, Duga S, Piccolo V, Ostuni R, Natoli G, Bronte V, Balzac F, Turco E, Hirsch E, Colombo MP, Sica A. Cancer Res. 2020 Jul 1;80(13):2874-2888. doi: 10.1158/0008-5472.CAN-19-2843. Epub 2020 Apr 7. - PMID: 18467122
Macrophage polarization in tumour progression.
Sica A, Larghi P, Mancino A, Rubino L, Porta C, Totaro MG, Rimoldi M, Biswas SK, Allavena P, Mantovani A. Semin Cancer Biol. 2008 Oct;18(5):349-55. doi: 10.1016/j.semcancer.2008.03.004. Epub 2008 Mar 26. Review. - PMID: 21995571
Macrophages in cancer and infectious diseases: the 'good' and the 'bad'.
Porta C, Riboldi E, Totaro MG, Strauss L, Sica A, Mantovani A. Immunotherapy. 2011 Oct;3(10):1185-202. doi: 10.2217/imt.11.116. Review. - PMID: 26267538
RORC1 Regulates Tumor-Promoting "Emergency" Granulo-Monocytopoiesis.
Sica A, Larghi P, Mancino A, Rubino L, Porta C, Totaro MG, Rimoldi M, Biswas SK, Allavena P, Mantovani A. Semin Cancer Biol. 2008 Oct;18(5):349-55. doi: 10.1016/j.semcancer.2008.03.004. Epub 2008 Mar 26. Review. - PMID:19616341
Cellular and molecular pathways linking inflammation and cancer.
Porta C, Larghi P, Rimoldi M, Totaro MG, Allavena P, Mantovani A, Sica A. Immunobiology. 2009;214(9-10):761-77. doi: 10.1016/j.imbio.2009.06.014. Epub 2009 Jul 17. Review. - PMID: 21948460
Origin and Functions of Tumor-Associated Myeloid Cells (TAMCs).
Sica A, Porta C, Morlacchi S, Banfi S, Strauss L, Rimoldi M, Totaro MG, Riboldi E. Cancer Microenviron. 2012 Aug;5(2):133-49. doi: 10.1007/s12307-011-0091-6. Epub 2011 Sep 24. Free PMC article. - PMID: 23049782
The p50 subunit of NF-κB orchestrates dendritic cell lifespan and activation of adaptive immunity.
Larghi P, Porta C, Riboldi E, Totaro MG, Carraro L, Orabona C, Sica A. PLoS One. 2012;7(9):e45279. doi: 10.1371/journal.pone.0045279. Epub 2012 Sep 25. Free PMC article. - PMID: 17825402
The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing.
De Santa F, Totaro MG, Prosperini E, Notarbartolo S, Testa G, Natoli G. Cell. 2007 Sep 21;130(6):1083-94. doi: 10.1016/j.cell.2007.08.019. Epub 2007 Sep 6. Free article. - PMID: 22127978
Enhanced interferon regulatory factor 3 binding to the interleukin-23p19 promoter correlates with enhanced interleukin-23 expression in systemic lupus erythematosus.
Smith S, Gabhann JN, Higgs R, Stacey K, Wahren-Herlenius M, Espinosa A, Totaro MG, Sica A, Ball E, Bell A, Johnston J, Browne P, O'Neill L, Kearns G, Jefferies CA. Arthritis Rheum. 2012 May;64(5):1601-9. doi: 10.1002/art.33494. Free article. - PMID: 18716661
The histone H3 lysine 27-specific demethylase Jmjd3 is required for neural commitment.
Burgold T, Spreafico F, De Santa F, Totaro MG, Prosperini E, Natoli G, Testa G. PLoS One. 2008 Aug 21;3(8):e3034. doi: 10.1371/journal.pone.0003034. Free PMC article.
Gallery Flow Cytometry unit
Equipment

Maria Grazia Totaro
Maria Grazia obtained her Master's degree in Biology Applied to Biomedical Research from the University of Insubria in Varese. Her academic and professional training was enriched through experiences at prestigious research institutes such as the Institute for Research in Biomedicine (IRB) in Bellinzona and the European Institute of Oncology (IEO) and Humanitas Institute in Milan, where she collaborated with leading researchers in the field of immunology.
During her PhD at Humanitas Institute, her passion for immunology led her to explore the fascinating world of flow cytometry and its countless applications.
After earning her PhD, Maria Grazia applied her expertise by joining the Imaging Unit at IFOM, led by Dr. Dario Parazzoli, as a flow cytometry specialist. Over the course of 10 years, she honed her skills and significantly contributed to the growth and development of the unit's flow cytometry activities, becoming a key reference figure.
In October 2024, she was appointed Head of Facility of the Flow Cytometry Unit, established following the division of the Imaging Unit, taking on the responsibility of leading this new entity towards scientific excellence.