Electron Microscopy Cells & Tissues

Unit info
Electron Microscopy (EM) is a powerful tool in cancer research, offering unparalleled resolution to observe structures that remain invisible under traditional light microscopes. Understanding cancer at the ultrastructural level is essential for studying tumor development, disease progression, and the effects of potential treatments.
At IFOM, our Electron Microscopy Cells & Tissues Core Facility provides expertise in electron microscopy to analyze a wide range of biological samples, from cultured cells to tissues. EM allows researchers to investigate the fine details of organelles and subcellular compartments, helping to uncover how oncogenes influence cellular organization and function. For example, EM enables the study of exosomes and their role in tumorigenesis, as well as the detection of morphological alterations induced by gene knockdown or knockout—changes that would otherwise go unnoticed.
Our facility offers a comprehensive range of advanced methods, including immuno-EM, correlative light-electron microscopy (CLEM), three-dimensional EM (3DEM), and quantitative ultrastructural analysis. By developing innovative techniques such as video CLEM for dynamic organelles and new sample preparation methods for 3DEM, we continuously refine our approach to meet the evolving needs of cancer researchers.
The Electron Microscopy Cells & Tissues Core Facility at IFOM provides a wide range of advanced imaging techniques to support research in cancer biology and cellular ultrastructure.
Current Techniques
- Transmission Electron Microscopy (TEM) – High-resolution imaging of cellular and subcellular structures.
- Scanning Transmission Electron Microscopy (STEM) – Combines TEM resolution with scanning capabilities for in-depth analysis.
- Immuno-Electron Microscopy (Immuno-EM) – Enables the localization of specific proteins within cells using gold nanoparticle labeling.
- Correlative Light and Electron Microscopy (CLEM) – Merges fluorescence microscopy with EM to link molecular and ultrastructural data.
- Three-Dimensional Electron Microscopy (3DEM) – Provides detailed 3D reconstructions of cellular structures.
Innovative Methods & Future Directions
We continuously develop and refine methodologies to enhance EM applications, including:
- Video-CLEM, ideal for studying fast-moving organelles.
- Discretized Rotator, a stereological tool for precise structural quantification.
- New sample preparation techniques for 3DEM, optimizing resolution and data accuracy.
Looking ahead, we aim to:
- Enhance CLEM and 3DEM methodologies for greater precision.
- Develop new quantitative analysis approaches using absolute stereological estimators.
- Design customized sample preparation protocols to meet the specific research needs of IFOM scientists.
Our ultimate goal is to establish modern guidelines for electron microscopy applications in oncology, ensuring that researchers have access to the most advanced and reliable imaging solutions.
Collaborations
The Facility is collaborating with IFOM scientists in the framework of the Athena strategic plan:
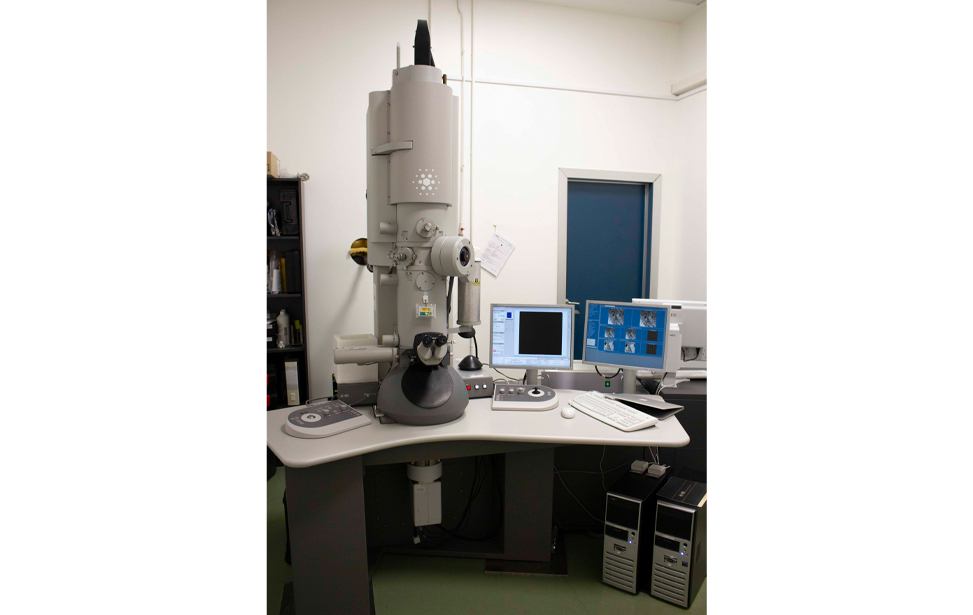
- Transmission electron microscope Tecnai G2 20 Twin - FEI Italia Srl (now Thermo Fisher Scientific). High-quality, high-resolution (S)TEM imaging and electron tomography.
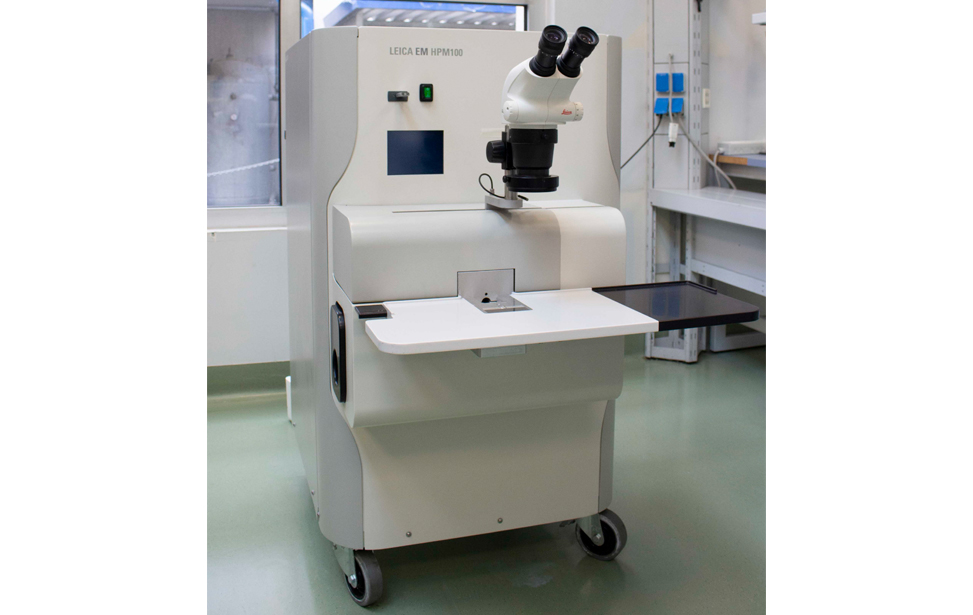
- Leica EM HPM100 is a high pressure freezing system for vitrifying samples up to 200μm in thickness. High Pressure Freezer for Cryofixation of Biological and Industrial Samples.
- Leica EM AFS2 performs freeze substitution and progressive lowering of temperature (PLT) techniques and allows low temperature embedding and polymerization of resins.
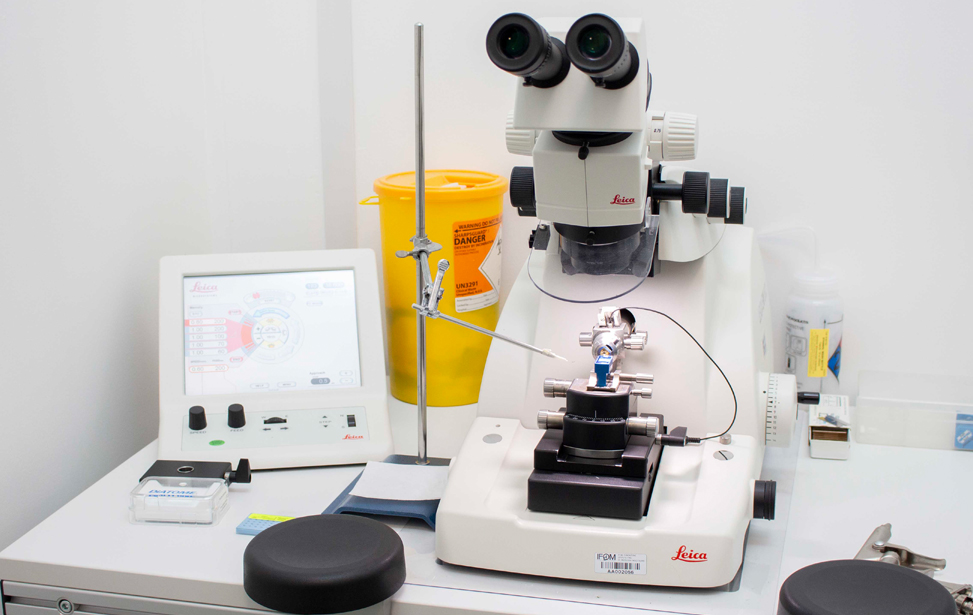
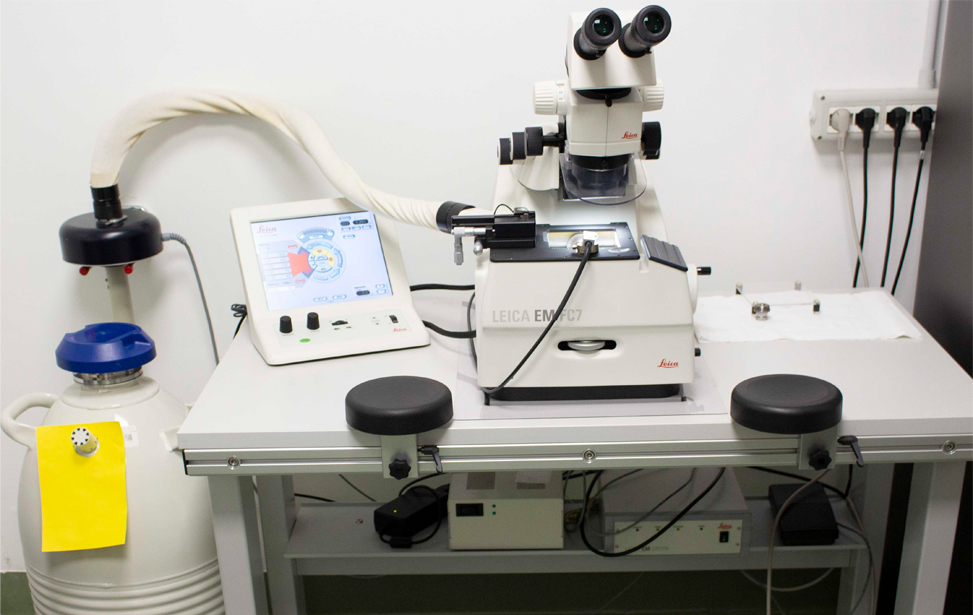
- Leica EM UC7 (at RT) consistent high-quality ultra- or semi-thin sections at room temperature
- Leica EM UC7 (CRYO-) Cryo-ultramicrotomy involves cutting ultra-thin sections from a frozen sample. These are thin enough to be viewed in a cryo-TEM. A cryo-ultramicrotome maintains temperatures below -140°C and can cut cryo-sections below 100nm in thickness.
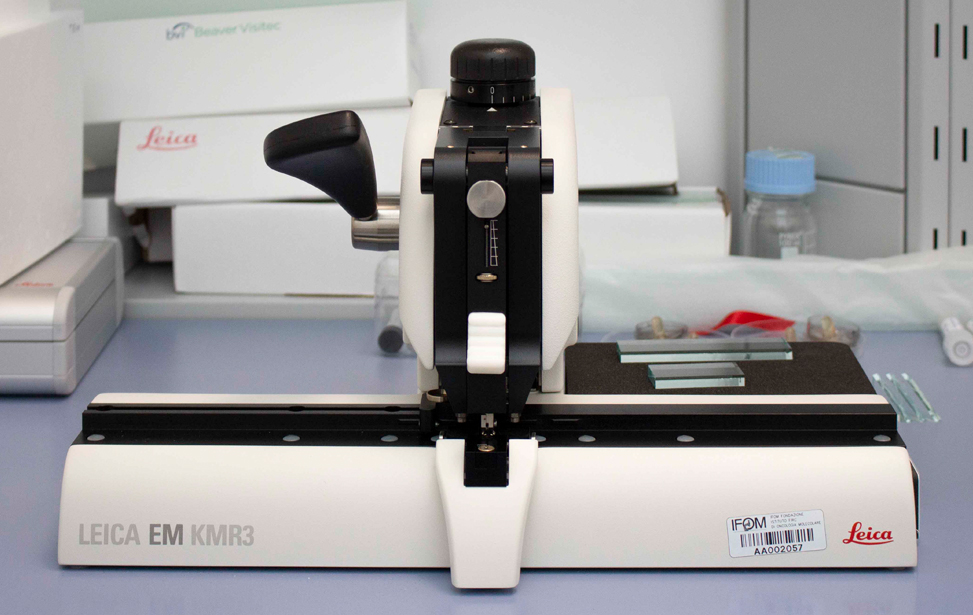
- Leica EM KMR 3 The balanced break method of the Leica EM KMR3 ensures perfect glass knives in three thicknesses 6.4 mm, 8 mm and 10 mm.
Facility Acknowledgment
According to our Core Facilities Guidelines, if Facility services were used in work described in a publication, the Facility must be acknowledged, citing both the Facillity name and its Research Resource Identifier (RRID). A typically acknowledgement might look like:
The research described was supported by the IFOM Electron Microscopy Cells & Tissues Core Facility (RRID: SCR_026863)
Co‑Authorship
If a staff member of an IFOM or Cogentech Core Facility contributes substantially to a published work, they should be included as a co‑author.
More information is available in Recommended guidelines and Acknowledging and citing core facilities
Examples of substantial contributions that may justify co‑authorship include:
- Contribution to experimental design or optimization
- Development or significant adaptation of methods or protocols
- Advanced data analysis and interpretation
- Intellectual input influencing the scientific direction of the study
Routine technical support or standard service provision generally warrants acknowledgment rather than co‑authorship.
Key Publications
- Martin S, Scorzoni S, Cordone S, Mazzagatti A, Beznoussenko GV, Gunn AL, Di Bona M, Eliezer Y, Leor G, Ben-Yishay T, Loffreda A, Cancila V, Rainone MC, Ippolito MR, Martis V, Bedin F, Garrè M, Vaites LP, Vasapolli P, Polo S, Parazzoli D, Tripodo C, Mironov AA, Cuomo A, Ben-David U, Bakhoum SF, Hatch EM, Ly P, Santaguida S.
A p62-dependent rheostat dictates micronuclei catastrophe and chromosome rearrangements.
Science. 2024 Aug 30;385(6712): eadj7446. doi: 10.1126/ science. adj 7446. Epub 2024 Aug 30. PMID: 39208097 - Kakogiannos N, Scalise AA, Martini E, Maderna C, Benvenuto AF, D'Antonio M, Carmignani L, Magni S, Gullotta GS, Lampugnani MG, Iannelli F, Beznoussenko GV, Mironov AA, Cerutti C, Bentley K, Philippides A, Zanardi F, Bacigaluppi M, Sigismund S, Bassani C, Farina C, Martino G, De Giovanni M, Dejana E, Iannacone M, Inverso D, Giannotta M.
GPR126 is a specifier of blood-brain barrier formation in the mouse central nervous system.
J Clin Invest. 2024 Jun 6;134(15): e165368. doi: 10.1172/JCI165368. PMID: 39087467. - Popęda M, Kowalski K, Wenta T, Beznoussenko GV, Rychłowski M, Mironov A, Lavagnino Z, Barozzi S, Richert J, Bertolio R, Myszczyński K, Szade J, Bieńkowski M, Miszewski K, Matuszewski M, Żaczek AJ, Braga L, Del Sal G, Bednarz-Knoll N, Maiuri P, Nastały P.
Emerin mislocalization during chromatin bridge resolution can drive prostate cancer cell invasiveness in a collagen-rich microenvironment.
Exp Mol Med. 2024 Sep;56(9):2016-2032. doi: 10.1038/s12276-024-01308-w. Epub 2024 Sep 2. PMID: 39218980 - Bastianello G, Porcella G, Beznoussenko GV, Kidiyoor G, Ascione F, Li Q, Cattaneo A, Matafora V, Disanza A, Quarto M, Mironov AA, Oldani A, Barozzi S, Bachi A, Costanzo V, Scita G, Foiani M.
Cell stretching activates an ATM mechano-transduction pathway that remodels cytoskeleton and chromatin.
Cell Rep. 2023 - Poli A, Pennacchio FA, Ghisleni A, di Gennaro M, Lecacheur M, Nastały P, Crestani M, Pramotton FM, Iannelli F, Beznusenko G, Mironov AA, Panzetta V, Fusco S, Sheth B, Poulikakos D, Ferrari A, Gauthier N, Netti PA, Divecha N, Maiuri P.
PIP4K2B is mechanoresponsive and controls heterochromatin-driven nuclear softening through UHRF1.
Nat Commun. 2023 Mar 14;14(1):1432. doi: 10.1038/s41467-023-37064-0.PMID: 36918565 - Frittoli E, Palamidessi A, Iannelli F, Zanardi F, Villa S, Barzaghi L, Abdo H, Cancila V, Beznoussenko GV, Della Chiara G, Pagani M, Malinverno C, Bhattacharya D, Pisati F, Yu W, Galimberti V, Bonizzi G, Martini E, Mironov AA, Gioia U, Ascione F, Li Q, Havas K, Magni S, Lavagnino Z, Pennacchio FA, Maiuri P, Caponi S, Mattarelli M, Martino S, d'Adda di Fagagna F, Rossi C, Lucioni M, Tancredi R, Pedrazzoli P, Vecchione A, Petrini C, Ferrari F, Lanzuolo C, Bertalot G, Nader G, Foiani M, Piel M, Cerbino R, Giavazzi F, Tripodo C, Scita G.
Tissue fluidification promotes a cGAS-STING cytosolic DNA response in invasive breast cancer.
Nat Mater. 2022 Dec 29. doi: 10.1038/s41563-022-01431-x. PMID: 36581770 - Kidiyoor GR, Li Q, Bastianello G, Bruhn C, Giovannetti I, Mohamood A, Beznoussenko GV, Mironov A, Raab M, Piel M, Restuccia U, Matafora V, Bachi A, Barozzi S, Parazzoli D, Frittoli E, Palamidessi A, Panciera T, Piccolo S, Scita G, Maiuri P, Havas KM, Zhou ZW, Kumar A, Bartek J, Wang ZQ, Foiani M.
ATR is essential for preservation of cell mechanics and nuclear integrity during interstitial migration.
Nat Commun. 2020 Sep 24;11(1):4828. doi: 10.1038/s41467-020-18580-9.PMID: 32973141 - Capaci V, Bascetta L, Fantuz M, Beznoussenko GV, Sommaggio R, Cancila V, Bisso A, Campaner E, Mironov AA, Wiśniewski JR, Ulloa Severino L, Scaini D, Bossi F, Lees J, Alon N, Brunga L, Malkin D, Piazza S, Collavin L, Rosato A, Bicciato S, Tripodo C, Mantovani F, Del Sal G.
Mutant p53 induces Golgi tubulo-vesiculation driving a prometastatic secretome.
Nat Commun. 2020 Aug 7;11(1):3945. doi: 10.1038/s41467-020-17596-5. PMID: 32770028 - Bisi S, Marchesi S, Rizvi A, Carra D, Beznoussenko GV, Ferrara I, Deflorian G, Mironov A, Bertalot G, Pisati F, Oldani A, Cattaneo A, Saberamoli G, Pece S, Viale G, Bachi A, Tripodo C, Scita G, Disanza A.
IRSp53 controls plasma membrane shape and polarized transport at the nascent lumen in epithelial tubules.
Nat Commun. 2020 Jul 14;11(1):3516. doi: 10.1038/s41467-020-17091-x. PMID: 32665580 - Kakogiannos N, Ferrari L, Giampietro C, Scalise AA, Maderna C, Ravà M, Taddei A, Lampugnani MG, Pisati F, Malinverno M, Martini E, Costa I, Lupia M, Cavallaro U, Beznoussenko GV, Mironov AA, Fernandes B, Rudini N, Dejana E, Giannotta M.
JAM-A Acts via C/EBP-α to Promote Claudin-5 Expression and Enhance Endothelial Barrier Function.
Circ Res. 2020 Sep 25;127(8):1056-1073. doi: 10.1161/CIRCRESAHA.120.316742. Epub 2020 Jul 15. PMID: 32673519 - Paulina Nastały, Divya Purushothaman, Stefano Marchesi, Alessandro Poli, Tobias Lendenmann, Gururaj Rao Kidiyoor, Galina V. Beznoussenko, Stefania Lavore, Orso Maria Romano, Dimos Poulikakos, Marco Cosentino Lagomarsino, Alexander A. Mironov, Aldo Ferrari, Paolo Maiuri.
Role of the nuclear membrane protein Emerin in front-rear polarity of the nucleus.
Nat Commun. 2020 May 1;11(1):2122. doi: 10.1038/s41467-020-15910-9.PMID: 32358486 - Palamidessi A, Malinverno C, Frittoli E, Corallino S, Barbieri E, Sigismund S, Beznoussenko GV, Martini E, Garre M, Ferrara I, Tripodo C, Ascione F, Cavalcanti-Adam EA, Li Q, Di Fiore PP, Parazzoli D, Giavazzi F, Cerbino R, Scita G.
Unjamming overcomes kinetic and proliferation arrest in terminally differentiated cells and promotes collective motility of carcinoma.
Nat Mater. 2019 Nov;18(11):1252-1263. - Yang JS, Hsu JW, Park SY, Li J, Oldham WM, Beznoussenko GV, Mironov AA, Loscalzo J, Hsu VW.
GAPDH inhibits intracellular pathways during starvation for cellular energy homeostasis.
Nature. 2018 Sep;561(7722):263-267. doi: 10.1038/s41586-018-0475-6. Epub 2018 Sep 12. - Malinverno C, Corallino S, Giavazzi F, Bergert M, Li Q, Leoni M, Disanza A, Frittoli E, Oldani A, Martini E, Lendenmann T, Deflorian G, Beznoussenko GV, Poulikakos D, Haur OK, Uroz M, Trepat X, Parazzoli D, Maiuri P, Yu W, Ferrari A, Cerbino R, Scita G.
Endocytic reawakening of motility in jammed epithelia.
Nat Mater. 2017 May;16(5):587-596. doi: 10.1038/nmat4848. Epub 2017 Jan 30. - Beznoussenko GV, Parashuraman S, Rizzo R, Polishchuk R, Martella O, Di Giandomenico D, Fusella A, Spaar A, Sallese M, Capestrano MG, Pavelka M, Vos MR, Rikers YG, Helms V, Mironov AA, Luini A.
Transport of soluble proteins the Golgi occurs by diffusion via continuities across cisternae.
Elife. 2014 May 27;3. doi: 10.7554/eLife.02009. - Kumar A, Mazzanti M, Mistrik M, Kosar M, Beznoussenko GV, Mironov AA, Garrè M, Parazzoli D, Shivashankar GV, Scita G, Bartek J, Foiani M.
ATR mediates a checkpoint at the nuclear envelope in response to mechanical stress.
Cell. 2014 Jul 31;158(3):633-46. doi: 10.1016/j.cell.2014.05.046. - Venditti R, Scanu T, Santoro M, Di Tullio G, Spaar A, Gaibisso R, Beznoussenko GV, Mironov AA, Mironov A Jr, Zelante L, Piemontese MR, Notarangelo A, Malhotra V, Vertel BM, Wilson C, De Matteis MA.
Sedlin controls the ER export of procollagen by regulating the Sar1 cycle.
Science. 2012 Sep 28;337(6102):1668-72. doi: 10.1126/science.1224947.
Methodological papers:
- Beznoussenko GV, and Mironov AA. Correlative video-light-electron microscopy of mobile organelles.
Methods Mol Biol. 2015; 1270: 321-46. doi: 10.1007/978-1-4939-2309-0_23. - Beznoussenko GV, Pilyugin SS, Geerts WJ, Kozlov MM, Burger KN, Luini A, Derganc J, Mironov AA.
Trans-membrane area asymmetry controls the shape of cellular organelles. Int J Mol Sci. 2015 Mar 9;16(3):5299-333. doi: 10.3390/ijms16035299. - Beznoussenko GV, Ragnini-Wilson A, Wilson C, Mironov AA.
Three-dimensional and immune electron microscopic analysis of the secretory pathway in Saccharomyces cerevisiae. Histochem Cell Biol. 2016 Nov;146(5):515-527. Epub 2016 Sep 3. - Mironov AA. Beznoussenko GV, Algorithm for Modern Electron Microscopic Examination of the Golgi Complex.
Methods Mol Biol. 2023;2557: 161-209. doi: 10.1007/978-1-0716-2639-9_12.PMID: 36512216 Review.
Photogallery
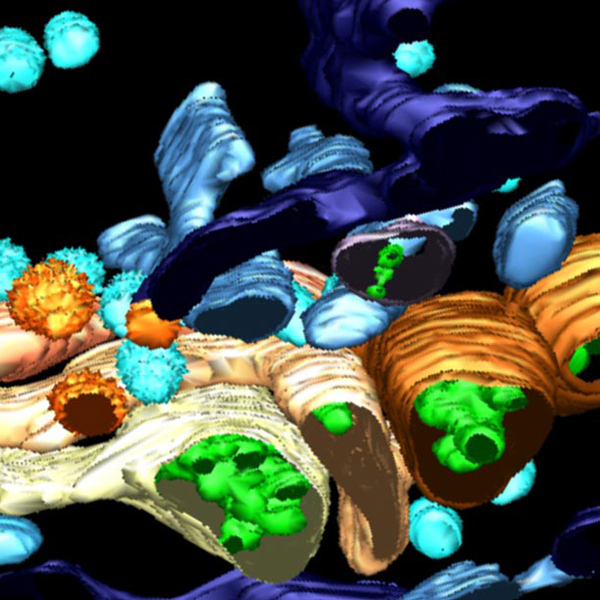
 3D reconstruction of Golgi apparatus
3D reconstruction of Golgi apparatus 3D reconstruction of Golgi apparatus
3D reconstruction of Golgi apparatus 3D reconstruction of Golgi apparatus
3D reconstruction of Golgi apparatusElectron Microscopy Instruments

Galina Beznusenko, M.D., PhD
Galina Beznusenko is the Head of the Electron Microscopy Cells & Tissues Core Facility at IFOM, where she has been a key figure since 2010. She graduated in Medicine and specialized in Obstetrics and Gynecology (1994, Ivanovo State Medical Institute, Russia) and Pathological Morphology (1996, Medical Academy of Postgraduate Training, St. Petersburg, Russia). Galina went on to pursue a PhD at Pirogov Russian National Research Medical University, focusing on Obstetrics and Gynecology, Histology, and Embryology.
In 2007, she earned the degree of Doctor of Science at the same university, specializing in Histology and Cell Biology. Galina has also held leadership roles, including Head of the Pathology Department at the Vladimir Medical Centre of Oncology for 5 years.
In 1997, Galina Beznusenko moved to Italy, where she worked at the Consorzio Mario Negri Sud for 13 years. In 2010, she and Alexandre Mironov established the Electron Microscopy Lab and EM Core Facility at IFOM, where they built a completely new and state-of-the-art facility that supports cutting-edge research in various scientific domains.